The Royal Swedish Academy of Sciences has decided to award Jacques Dubochet, Joachim Frank, and Richard Henderson the Nobel Prize for Chemistry 2017 for "developing cryo-electron microscopy for the high-resolution structure determination of biomolecules in solution". Scientific Background on the Nobel Prize in Chemistry 2017: The Development of Cryo-Electron Microscopy
Cryo-EM is being led by the technology and innovation behind Gatan's direct detection cameras (K3® and K2®).
The efficient frontier of resolution Advantages Workflow for cryo-EM
What is single-particle cryo-EM?
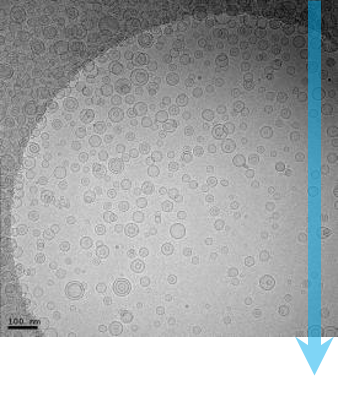
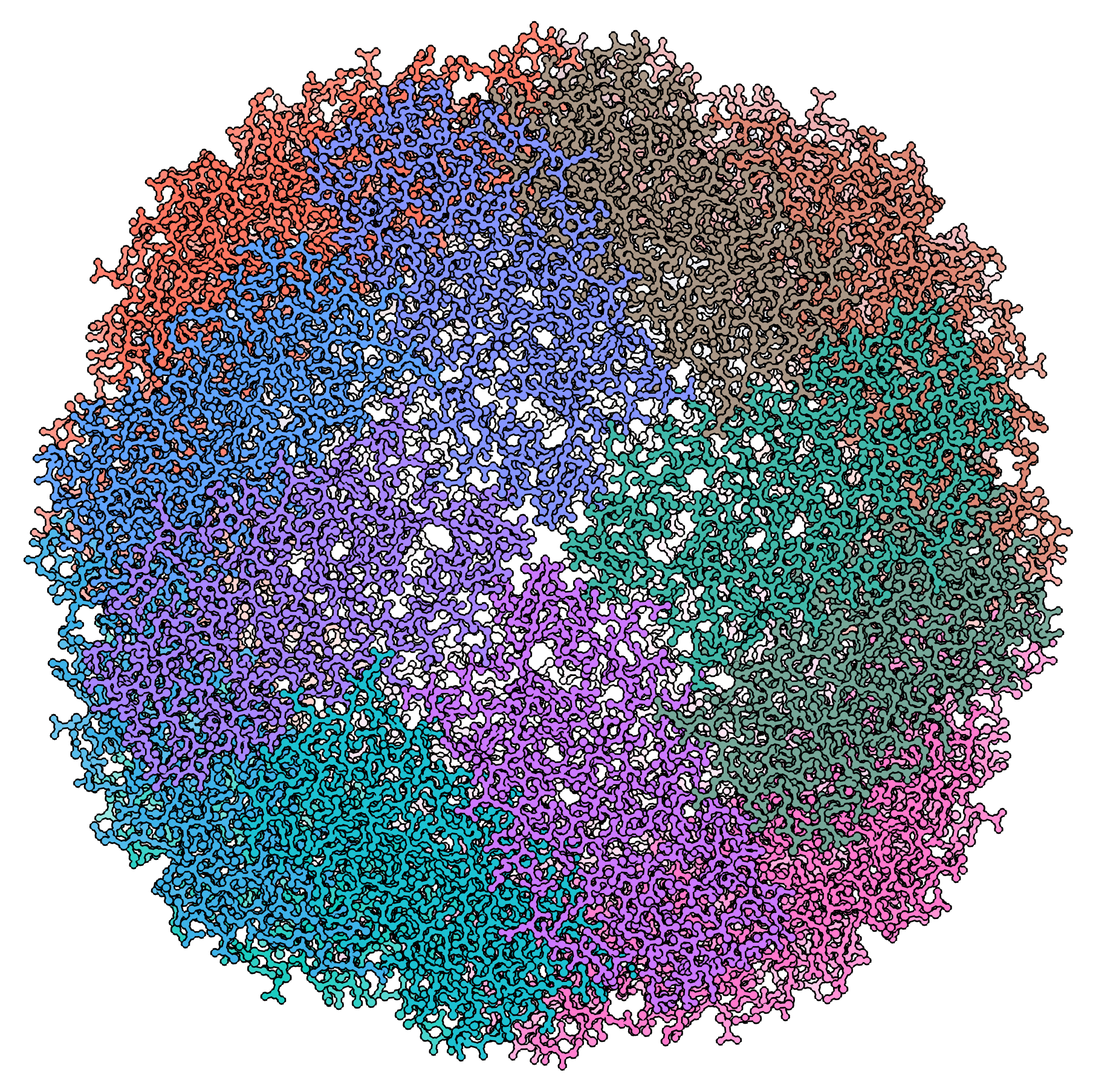 Single-particle cryo-electron microscopy (cryo-EM), is an increasingly popular technique used by structural biologists to solve structures at atomic resolution. This technique complements x-ray crystallography because it reveals structural details without the need for a crystalline specimen. Through the examination of a frozen-hydrated specimen in vitreous (non-crystalline) ice, the specimen ultrastructure, buffer composition, and ligand distribution from its native state are maintained. Cryo-EM also complements structural studies using nuclear magnetic resonance (NMR) in that it enables the study of specimens larger than 90 kDa. Structural biologists frequently use cryo-EM to study viruses, small organelles, and macromolecular biological complexes, purified proteins as well as molecular interactions in supramolecular assemblies or machines.
Single-particle cryo-electron microscopy (cryo-EM), is an increasingly popular technique used by structural biologists to solve structures at atomic resolution. This technique complements x-ray crystallography because it reveals structural details without the need for a crystalline specimen. Through the examination of a frozen-hydrated specimen in vitreous (non-crystalline) ice, the specimen ultrastructure, buffer composition, and ligand distribution from its native state are maintained. Cryo-EM also complements structural studies using nuclear magnetic resonance (NMR) in that it enables the study of specimens larger than 90 kDa. Structural biologists frequently use cryo-EM to study viruses, small organelles, and macromolecular biological complexes, purified proteins as well as molecular interactions in supramolecular assemblies or machines.
During single-particle cryo-EM, a transmission electron microscope (TEM) is used to record high-resolution images of thousands to hundreds of thousands of identical, but randomly oriented, particles (molecules) from each specimen. These images are then grouped, aligned, and averaged with image classification algorithms to distinguish between multiple orientations of the 3D molecule. With a well-behaved sample, cryo-EM can solve molecular structures with a resolution below 1.5 Å; a resolution level that was inconceivable only a few years ago.
This rapid improvement of cryo-EM resolution has been called the “resolution revolution” and is the direct result of direct detection cameras. Conventional cameras for electron microscopy use a scintillator to convert the electron image to a light image, and a fiber optic faceplate to transfer the image to a CCD or CMOS image sensor for analog recording. When the results are converted and recorded, there is a loss of high-resolution detail that has long held cryo-EM from achieving its true potential.
Direct detection cameras now directly measure the electron image to avoid the loss of detail from the conventional camera conversion steps. The Gatan K3 camera is a unique direct detection camera that uses an electron counting with super-resolution technology to record the image. This technology recognizes and counts individual image electrons in real-time, while it rejects the analog read noise. As shown with Gatan's direct detection cameras, the detective quantum efficiency (DQE) performance at half Nyquist is unparalleled.
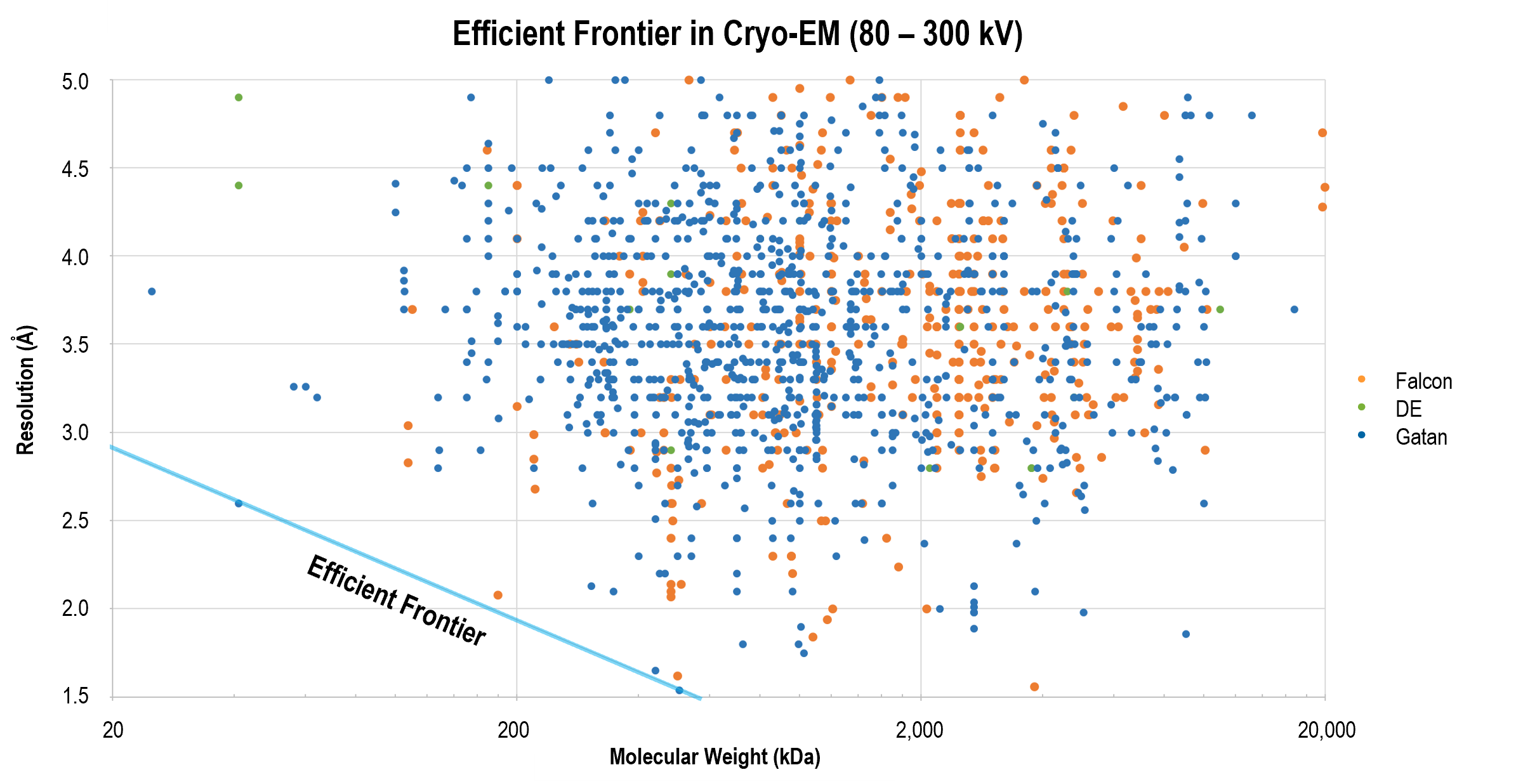
The efficient frontier of resolution
Gatan's direct detection camera, often in combination with the GIF Quantum LS imaging filter, continues to yield breakthrough results that define the efficient frontier of resolution. Cutting edge publications show these products along with improved data acquisition and image processing strategies remain at the forefront of higher resolution reconstructions for smaller molecules, such as the 2.2 Å structure of β-galactosidase.
The graph above is a comparison between the resolution and molecule size for published single-particle cryo-EM structures. Gatan's direct detectors have been employed in all the structures which define the high-resolution frontier across a range of molecular sizes.
Advantages of single-particle cryo-EM
Now that single-particle cryo-EM provides structures with comparable resolution to x-ray crystallography, there are a number of unique advantages that are helping the technique gain traction amongst structural biologists:
| Capability | Advantage |
|---|---|
| Examines structures in their native, hydrated state | Maintains your specimen in a biologically relevant environment, including sample concentration and buffer composition |
| Allows study of larger assemblies | Useful to characterize molecules larger than 150 kDa that include multiple subunits that are heterogeneous, metastable or extremely difficult to crystallize |
| Elucidates atomic resolution structures | Enables observation of asymmetric side chains, hydrogen bonds, and water molecules in addition to alpha helices and beta sheets |
| Controls the chemical environment | Allows you to vary experimental conditions to examine molecules in different functional states |
| Eliminates crystallization steps | Avoids long, uncertain preparation steps; shortens your time to publish |
Workflow for single-particle cryo-EM
|
|
Step 1: Purify To study molecules by single-particle cryo-EM, a specimen must be purified and structurally intact to produce high-quality 3D reconstructions. Ideally, you need to maintain the specimen in a buffer solution that keeps it biochemically active. Molecules within the specimen should be in a high enough concentration that you can study them under a microscope, but not so high that they aggregate together. Finally, experimental conditions should be optimized to promote a uniform conformational state for your molecule of interest. |
|
|
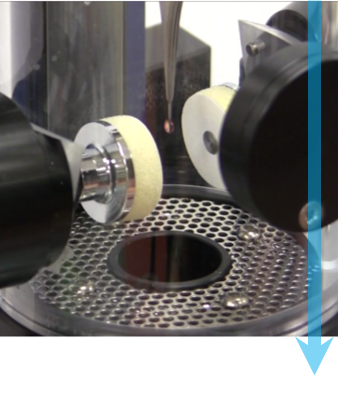
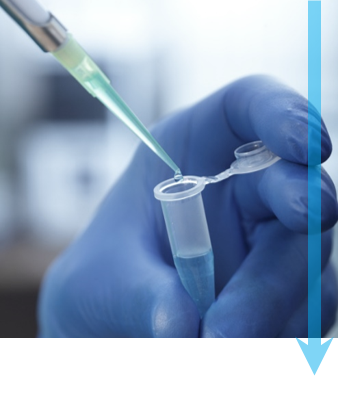
Step 2: Plunge freeze Each specimen is frozen to prevent it from freeze-drying within the vacuum of the microscope. Nearly instantaneous freezing prevents the formation of ice crystals that will disrupt the structure of the specimen. To start, you apply a small amount of the specimen in solution to a TEM grid before the blotting paper is briefly used to remove excess liquid. The TEM grid is then plunged into liquid ethane or a mixture of ethane/propane to quickly wet the specimen, remove heat, and create non-crystalline or vitreous ice. The image shows the Cryoplunge® 3 system just prior to blotting and submersion of the specimen into the liquid cryogen. |
|
|
Step 3: Transfer to TEM Once frozen, you transfer the specimen to a specialized TEM holder that keeps it at liquid nitrogen temperature. To prevent specimen contamination, a cryo-workstation protects the specimen when you load it into the holder, then a cryo-shield encapsulates it during the transfer from the workstation to the TEM. As seen here, the cryo-transfer holder is being retracted from the workstation prior to insertion into the TEM. |
|
|
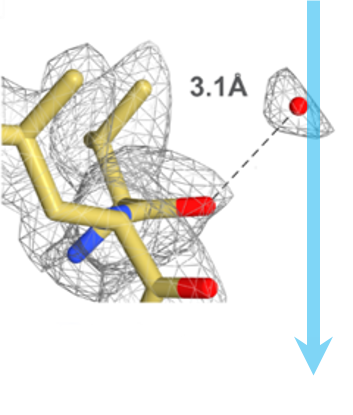
Step 4: Image specimen A specimen’s structural integrity is damaged when exposed to electrons, and typically a total dose of 10 – 30 e-/Å2 can be used before high-resolution structural information is lost. To prevent specimen damage, low-dose imaging procedures are used to navigate to the desired areas and focus the electron beam before acquiring images. The high DQE associated with Gatan's direct detector electron counting and super-resolution modes enables you to acquire the highest quality images of delicate biological samples. This high image signal-to-noise ratio allows you to discern water molecules, ions, and ligand structures in the 3D particle reconstruction. You can further improve image quality by utilizing the dose fractionation feature on Gatan's direct detection cameras, which saves full frames at up to 75 frames per second, to later correct for specimen motion and minimize drift. |
|
|
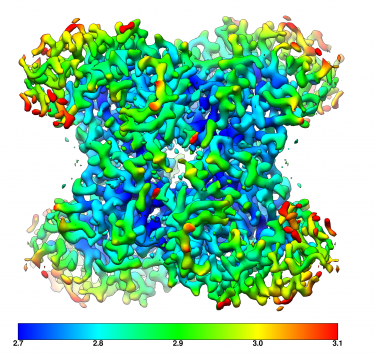
Step 5: Analyze and reconstruct Once imaged, the Gatan Microscopy Suite® software supports your analysis and export of data into multiple formats. Data from Gatan cameras is imported into a wide variety of 3rd party software tools for 3D reconstruction and visualization, including EMAN, Frealign, Relion, and many others. The image shows a 3D cryo-EM density of 20S proteasome at 2.8 Å resolution. |
Optimized for cryo-EM and cryo-ET to gain further insight into system function and disease progression at the molecular level.
The next generation cryo-holder for your high-resolution cryo-EM and tomography experiments.
Sets a new standard for the efficient, high-throughput collection of low-dose, single-particle, cryo-EM datasets from Gatan’s cameras.
The next-generation plasma tool to remove hydrocarbon contamination from TEM and SEM samples and holders.
The best-in-class imaging filter for cryo-EM is now available with EELS and EFTEM.
DigitalMicrograph, also known as Gatan Microscopy Suite, drives your digital cameras and surrounding components to support key applications including tomography, in-situ, spectrum and diffraction imaging, plus more.
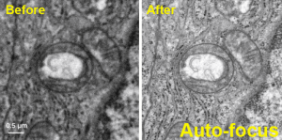
Facilitate your HREM assays by automatically adjusting the critical imaging parameters of a TEM microscope focus, stigmation, and beam tilt.
-
 Transthyretin resolved to 4.6 Å at 100 keV
Transthyretin resolved to 4.6 Å at 100 keV
-
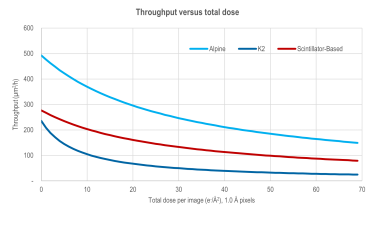 Throughput versus total dose comparison - Alpine
Throughput versus total dose comparison - Alpine
-
 Aldolase resolved to 3.07 Å at 100 keV
Aldolase resolved to 3.07 Å at 100 keV
-
 Apoferritin resolved to 2.7 Å at 100 keV
Apoferritin resolved to 2.7 Å at 100 keV
-
 Imaging molecules in their native environment: Cryo-electron tomography of PCDH15 complexes in mouse stereocilia
Imaging molecules in their native environment: Cryo-electron tomography of PCDH15 complexes in mouse stereocilia
-

 Context matters: Cryo-ET reveals the impact of the cellular environment on nuclear pore complex
Context matters: Cryo-ET reveals the impact of the cellular environment on nuclear pore complex
-
 Cryo EM reveals mechanisms of gating and drug modulation in 5 HT3A receptors webinar
Cryo EM reveals mechanisms of gating and drug modulation in 5 HT3A receptors webinar
-
 Webinar: High-resolution with the CryoARM/K3 combo: SerialEM, Latitude, and future data collection
Webinar: High-resolution with the CryoARM/K3 combo: SerialEM, Latitude, and future data collection
-

 Using Structural Biology to Drive Pandemic Preparedness Webinar
Using Structural Biology to Drive Pandemic Preparedness Webinar
-
 Elsa Cryo-Transfer Workstation
Elsa Cryo-Transfer Workstation
In situ structural analysis of SARS-CoV-2 spike reveals flexibility mediated by three hinges
Turoňová, B.; Sikora, M.; Schürmann, C.; Hagen, W. J. H.; Welsch, S.; Blanc, F. E. C.; von Bülow, S.; Gecht, M.; Bagola, K.; Hörner, C.; van Zandbergen, G.; Landry, J.; de Azevedo, N. T. D.; Mosalaganti, S.; Schwarz, A.; Covino, R.; Mühlebach, M. D.; Hummer, G.; Locker, J. K.; Beck, M.
Molecular architecture of the SARS-CoV-2 virus
Yao, H.; Song, Y.; Chen, Y.; Wu, N.; Xu, Sun, C.; Zhang, J.; Weng, T.; Zhang, Z.; Wu, Z.; Cheng, L,; Shi, D.; Lu, X.; Lei, J.; Crispin, M.; Shi, Y.; Li, L.; Li, S.
Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation
Wrapp, D.; Wang, N.; Corbett, K. S.; Goldsmith, J. A.; Hsieh, C. -L.; Abiona, O.; Graham, B. S.; McLellan, J. S.
Cryo-EM structure of the Ebola virus nucleoprotein–RNA complex at 3.6 Å resolution
Sugita, Y.; Matsunami, H.; Kawaoka, Y.; Noda, T.; Wolf, M.
Venugopal, H.; Mobbs, J.; Taveneau, C.; Fox, D. R.; Vuckovic, Z.; Knott, G.; Grinter, R.; Thal, D.; Mick, S.; Czarnik, C.; Ramm, G.
Chan, L. M.; Courteau, B. J.; Maker, A.; Wu, M.; Basanta, B.; Mehmood, H.; Bulkley, D.; Joyce, D.; Lee, B. C.; Mick, S.; Gulati, S.; Lander, G. C.; Verba, K. A.
Electron-counting MicroED data with the K2 and K3 direct electron detectors
Clabbers, M. T. B.; Martynawycz, M. W.; Hattne, J.; Nannenga, B. L.; Gonen, T.
Yang, T.; Xua, H.; Zoua, X.
Time-resolved cryo-EM using a combination of droplet microfluidics with on-demand jetting
Torino, S.; Dhurandhar, M.; Stroobants, A.; Claessens, R.; Efremov, R. G.
Szemán, A. J. K.; Stráner, P.; Jákli, I.; Hosogi, N.; Harmat, V.; Menyhárd, D. K.; Perczel, A.
Structural basis of actin filament assembly and aging
Oosterheert, W.; Klink, B. U.; Belyy, A.; Pospich, S.; Raunser, S.
Helical ultrastructure of the metalloprotease meprin α in complex with a small molecule inhibitor
Bayly-Jones, C.; Lupton, C. J.; Fritz, C.; Venugopal, H.; Ramsbeck, D.; Wermann, M.; Jäger,C.; Marco, A. D.; Schilling, S.; Schlenzig, D.; Whisstock, J. C.
Stern, A. M.; Yang, Y.; Meunier, A. L.; Liu, W.; Cai, Y.; Ericsson, M.; Liu, L.; Goedert, M.; Scheres, S. H. W.; Selkoe, D.J
Cryo-EM structure of the human NKCC1 transporter reveals mechanisms of ion coupling and specificity
Neumann, C.; Rosenbæk, L. L.; Flygaard, R. K.; Habeck, M.; Karlsen, J. L.; Wang, Y.; Larsen, K. L.; Gad, H. H.; Hartmann, R.; Lyons, J. A; Fenton, R. A.; Nissen, P.
Algal photosystem I dimer and high-resolution model of PSI-plastocyanin complex
Naschberger, A.; Mosebach, L.; Tobiasson, V.; Kuhlgert, S.; Scholz, M.; Perez-Boerema, A.; Ho, T. T. H.; Vidal-Meireles, A.; Takahashi, Y.; Hippler, M.; Amunts, A.
Chang, M. R.; Tomasovic, L.; Kuzmina, N. A.; Ronk, A. J.; Byrne, P. O.; Johnson, R.; Storm, N.; Olmedillas, E.; Hou, Y. J.; Schäfer, A.; Leist, S. R.; Tse, L. T.; Ke, H.; Coherd, C.; Nguyen, K.; Kamkaew, M.; Honko, A.; Zhu, Q.; Alter, G.; Saphire, E. O.; McLellan, J. S.; Griffiths, A.; Baric, R. S.; Bukreyev. A.; Marasco. W. A.
Frieg, B.; Geraets, J. A.; Strohäker, T.; Dienemann, C.; Mavroeidi, P.; Jung, B. C.; Kim, W. S.; Lee, S. J.; Xilouri, M.; Zweckstetter. M.; Schröder. G. F.
Lyu, M.; Ayala, J. C.; Chirakos, I.; Su, C. -C.; Shafer, W. M.; Yu, E. W.
Organic crystal growth: Hierarchical self-assembly involving nonclassical and classical steps
Biran, I.; Rosenne, S.; Weissman, H.; Tsarfati, Y.; Houben, L.; Rybtchinski. B.
Structure of the PAPP-ABP5 complex reveals mechanism of substrate recognition
Judge, R. A.; Sridar, J.; Tunyasunvunakool, K.; Jain, R.; Wang, J. C. K.; Ouch, C.; Xu, J.; Mafi, A.; Nile, A. H.; Remarcik, C.; Smith, C. L.; Ghosh, C.; Xu, C.; Stoll, V.; Jumper, J.; Singh, A. H.; Eaton, D.; Hao, Q.
Structures of a phycobilisome in light-harvesting and photoprotected states
Domínguez-Martín, M. A.; Sauer, P. V.; Kirst, H.; Sutter, M.; Bína, D.; Greber, B. J.; Nogales, E. ;Polívka, T.; Kerfeld, C. A.
Zhu, J.; Huang, W.; Zhao, J., Huynh, L.; Taylor, D. J.; Harris, M. E.
Miyakawa, T.; Yang, J.; Kawasaki, M.; Adachi, N.; Fujii, A.; Miyauchi, Y.; Muramatsu, T.; Moriya, T.; Senda, T.; Tanokura, M.
Kuhle, B.; Hirschi, M.; Doerfel, L. K.; Lander, G. C.; Schimmel, P.
Zhao, J.; Makhija, S.; Zhou, C.; Zhang, H.; Wang, Y.; Muralidharan, M.; Huang, B.; Cheng, Y.
Mechanistic details of CRISPR-associated transposon recruitment and integration revealed by cryo-EM
Park, J. U.; Tsai, A. W. -T.; Chen, T. H.; Peters, J. E.; Kellogg, E. H.
Specific recognition and ubiquitination of slow-moving ribosomes by human CCR4-NOT
Absmeier, E.; Chandrasekaran, V.; O'Reilly, F. J.; Stowell, J. A. W.; Rappsilber, J.; Passmore, L. A.
Cryo-electron microscopy of extracellular vesicles
Cai, K.; Sibert, B. S.; Kumar, A.; Yang, J.; Larson, M.; Thompson, K.; Wright, E. R.
Zhang, Z.; Li, Y.; Zhou, W.; Li, Y.; Chiu, W.; Cui, Y.
Peck, J. V.; Strauss, J. D.; Fay, J. F.
Cryo-EM structure of an active bacterial TIR–STING filament complex
Morehouse, B. R.; Yip, M. C. J.; Keszei, A. F. A.; McNamara-Bordewick, N. K.; Shao, S.; Kranzusch, P. J.
Maintaining the momentum in cryoEM for biological discovery
Thompson, R.; Halfon, Y.; Aspinall, L.; White, J.; Hirst, I. J.; Wang, Y.; Darrow, M.; Muench, S. P.
Orris, B.; Huynh, K. W.; Ammirati, M.; Han, S.; Bolaños, B.; Carmody, J.; Petroski, M. D.; Bosbach, B.; Shields, D. J.; Stivers, J. T.
Structural analysis of the basal state of the Artemis:DNA-PKcs complex
Watanabe, G.; Lieber, M. R.; Williams, D. R.
Flipped over U: structural basis for dsRNA cleavage by the SARS-CoV-2 endoribonuclease
Frazier, M. N.; Wilson, I. M.; Krahn, J. M.; Butay, K. J.; Dillard, L. B.; Borgnia, M. J.; Stanley, R. E.
Cryo-EM structure of the HIV-1 Pol polyprotein provides insights into virion maturation
Harrison, J. J. E. K.; Passos, D. O.; Bruhn, J. F.; Baumanlynda Tuberty, J. D.; Destefano, J. J.; Ruiz, F. X.; Lyumkis, D.; Arnold, E.
Structures and gating mechanisms of human bestrophin anion channels
Owji, A. P.; Wang, J.; Kittredge, A.; Clark, Z.; Zhang, Y.; Hendrickson, W. A.; Yang, T.
High-resolution structure determination using high-throughput electron cryo-tomography
Liu, H. -F.; Ye Zhou, Y.; Bartesaghi, A.
Structural insights into dsRNA processing by Drosophila Dicer-2–Loqs-PD
Su, S.; Wang, J.; Deng, T.; Yuan, X.; He, J.; Liu, N.; Li, X.; Huang, Y.; Wang, H. -W.; Ma, J.
Structures and mechanism of the plant PIN-FORMED auxin transporter
Ung, K. L.; Winkler, M.; Schulz, L.; Kolb, M.; Janacek, D. P.; Dedic, E.; Stokes, D. L.; Hammes, U. Z.; Pedersen, B. P.
A peroxisomal ubiquitin ligase complex forms a retrotranslocation channel
Feng, P.; Wu, X.; Erramilli, S. K.; Paulo, J. A.; Knejski, P.; Gygi, S. P.; Kossiakoff, A. A.; Rapoport, T. A.
Cryo-EM structures of SARS-CoV-2 Omicron BA.2 spike
Stalls, V.; Lindenberger, J.; Gobeil, S. M. -C.; Henderson, R.; Parks, R.; Barr, M.; Deyton, M.; Martin, M.; Janowska, K.; Huang, X.; May, A.; Speakman, M.; Beaudoin, E.; Kraft, B.; Lu, X.; Edwards, R. J.; Eaton, A.; Montefiori, D. C.; Williams, W. B.; Saunders, K. O.; Wiehe, K.; Haynes, B. F.; Acharya, P.
Structure and flexibility of the yeast NuA4 histone acetyltransferase complex
Zukin, S. A.; Marunde, M. R.; Popova, I. K.; Nogales, E.; Patel, A. B.
Structure and flexibility of the yeast NuA4 histone acetyltransferase complex
Zukin, S. A.; Marunde, M. R.; Popova, I. K.; Nogales, E.; Patel, A. B.
Martinez-Molledo, M.; Nji, E.; Reyes, N.
Role of aIF5B in archaeal translation initiation
Kazan, R.; Bourgeois, G.; Lazennec-Schurdevin, C.; Larquet, E.; Mechulam, Y.; Coureux, P. -D.; Schmitt, E.
Ion complexation waves emerge at the curved interfaces of layered minerals
Whittaker, M. L.; Ren, D.; Ophus, C.; Zhang, Y.; Waller, L.; Gilbert, B.; Banfield, J. F.
Compact IF2 allows initiator tRNA accommodation into the P site and gates the ribosome to elongation
Basu, R. S.; Sherman, M. B.; Gagnon, M. G.
Structural Basis for pH-gating of the K+ channel TWIK1 at the selectivity filter
Turney, T. S.; Li, V.; Brohawn, S. G.
Cryo-EM structures of SARS-CoV-2 Omicron BA.2 spike
Stalls, V.; Lindenberger, J.; Gobeil, S. M. -C.; Henderson, R.; Parks, R.; Barr, M.; Deyton, M.; Martin, M.; Janowska, K.; Huang, X.; May, A.;l Speakman, M.; Beaudoin, E.; Kraft, B.; Lu, X.; Edwards, R. J.; Eaton, A.; Montefiori, D. C.; Williams, W.; Saunders, K. O.; Wiehe, K.; Haynes, B. F.; Acharya, P.
Xu, Y.; Margetts, M. B.; Venugopal, H.; Menting, J. G.; Kirk, N. S.; Croll, T. I.; Delaine, C.; Forbes, B. E.; Lawrence, M. C.
Structure of S1PR2–heterotrimeric G13 signaling complex
Chen, H.; Chen, K.; Huang, W.; Staudt, L. M.; Cyster, J. G.; Li, X.
Structure of the type V-C CRISPR-Cas effector enzyme
Kurihara, N.; Nakagawa, R.; Hirano, H.; Okazaki, S.; Tomita, A.; Kobayashi, K.; Kusakizako, T.; Nishizawa, T.; Yamashita, K.; Scott, D. A.; Nishimasu, H.; Nureki, O.
Cryogenic TEM imaging of artificial light harvesting complexes outside equilibrium
Krishnaswamy, S. R.; Gabrovski, I. A.; Patmanidis, I.; Stuart, M. C. A.; de Vries A. H.; Pshenichnikov, M. S.
Assembly mechanism of the pleomorphic immature poxvirus scaffold
Hyun, J.; Matsunami, H.; Kim, T. G.; Wolf, M.
The giant Mimivirus 1.2 Mb genome is elegantly organized into a 30 nm helical protein shield
Villalta, A.; Schmitt, A.; Estrozi, L. F.; Quemin, E. R. J.; Alempic, J. -M.; Lartigue, A.; Pražák, V.; Belmudes, L.; Vasishtan, D.; Colmant, A. M. G.; Honoré, F. A.; Couté, Y.; Grünewald, K.; Abergel, C.
Zhao, W.; Jensen, G. J.
Structural and functional impact by SARS-CoV-2 Omicron spike mutations
Zhang, J.; Cai, Y.; Lavine, C. L.; Peng, H.; Zhu, H.; Anand, K.; Tong, P.; Gautam, A.; Mayer, M. L.; Rits-Volloch, S.; Wang, S.; Sliz, P.; Wesemann, D. R.; Yang, W.; Seaman, M. S.; Lu, J.; Xiao, T.; Chen, B.
Capturing the swelling of solid-electrolyte interphase in lithium metal batteries
Zhang, Z.; Li, Y.; Xu, R.; Zhou, W.; Li, Y.; Oyakhire, S. T.; Wu, Y.; Xu, J.; Wang, H.; Yu, Z.; Boyle, D. T.; Huang, W.; Ye, Y.; Chen, H.; Wan, J.; Bao, Z.; Chiu, W.; Cui, Y.
Nuclear pores dilate and constrict in cellulo
Zimmerli, C. E.; Allegretti, M.; Rantos, V.; Goetz, S. K.; Obarska-Kosinska, A.; Zagoriy, I.; Halavatyi, A.; Hummer, G.; Mahamid, J.; Kosinski, J.; Beck, M.
Scaffolding protein CcmM directs multiprotein phase separation in β-carboxysome biogenesis
Zang, K.; Wang, H.; Hartl, F. U.; Hayer-Hartl, M.
Alegre, K. O.; Paknejad, N.; Su, M.; Lou, J. -S.; Huang, J.; Jordan, K. D.; Eng, E. T.; Meyerson, J. R.; Hite, R. K.; Huang, X. -Y.
Emmanuel, S. N.; Smith, J. K.; Hsi, J.; Tseng, Y. -S.; Kaplan, M.; Mietzsch, M.; Chipman, P.; Asokan, A.; Robert McKenna, R.; Agbandje-McKenna, M.
Yu, H.; Hamaguchi, T.; Nakajima, Y.; Kato, K.; Kawakami, K.; Akita, F.; Yonekura, K.; Shen, J. -R.
Peng, Q.; Peng, R.; Yuan, B.; Zhao, J.; Wang, M.; Wang, X.; Wang, Q.; Sun, Y.; Fan, Z.; Qi, J.; Gao, G. F.; Shi, Y.
A simple pressure-assisted method for MicroED specimen preparation
Zhao, J.; Xu, H.; Lebrette, H.; Carroni, M.; Taberman, H.; Hogbom, M.; Zou, X.
Structure-guided multivalent nanobodies block SARS-CoV-2 infection and suppress mutational escape
Koenig, P. -D.; Das, H.; Liu H.; Kümmerer, B. M.; Gohr, F. N.; Jenster, L. -M.; Schiffelers, L. D. J.; Tesfamariam, Y. M.; Uchima, M.; Wuerth, J. D.; Gatterdam, K.; Ruetalo, N.; Christensen, M. H.; Fandrey, C. I.; Normann, S.; Tödtmann, J.; M. P.; Pritzl, S.; Hanke, L.; Boos, J.; Yuan, M.; Zhu, X.; Schmid-Burgk, J. L.; Kato, H.; Schindler, M.; Wilson, I. A.; Geyer, M.; Ludwig, K. U.; Hällberg, M.; Wu, N. C.; Schmidt, F. I.
Ultrapotent antibodies against diverse and highly transmissible SARS-CoV-2 variants
Wang, L.; Zhou, T.; Zhang, Y.; Yang, E. S.; Schramm, C. A.
Cryo-EM structures of human coagulation factors V and Va
Ruben, E. A.; Rau, M. J.; Fitzpatrick, J. A. J.; Di Cera, E.
Wei, H.; Qian, X.; Xie, F.; Cui, D.
Native-like SARS-CoV-2 spike glycoprotein expressed by ChAdOx1 nCoV-19/AZD1222 vaccine
Watanabe, Y.; Mendonça, L.; Allen, E. R.; Howe, A.; Lee, M.; Allen, J. D.; Chawla, H.; Pulido, D.; Donnellan, F.; Davies, H.; Ulaszewska, M.; Belij-Rammerstorfer, S.; Morris, S.; Krebs, A. -S.; Dejnirattisai, W.; Mongkolsapaya, J.; Supasa, P.; Screaton, G. R.; Green, C. M.; Lambe, T.; Zhang, P.; Gilbert, S. C.; Crispin, M.
Simplified approach for preparing graphene oxide TEM grids for stained and vitrified biomolecules
Kumar, A.; Sengupta, N.; Dutta, S.
Shafiq, A.; Suwakulsiri, W.; Rai, A.; Chen, M.; Greening, D. W.; Zhu, H. -J.; Xu, R.; Simpson, R. J.
Structure of a microtubule-bound axonemal dynein
Walton, T.; Wu, H.; Brown, A.
Mechanism of SARS-CoV-2 polymerase stalling by remdesivir
Kokic, G.; Hillen, H. S.; Tegunov, D.; Dienemann, C.; Seitz, F.; Schmitzova, J.; Farnung, L.; Siewert, A.; Höbartner, C.; Cramer, P.
Stabilizing the closed SARS-CoV-2 spike trimer
Juraszek, J.; Rutten, L.; Blokland, S.; Bouchier, P.; Voorzaat, R.; Ritschel, T.; Bakkers, M. J. G.; Renault , L. L. R.; Langedijk, J. P. M.
Development and structural basis of a two-MAb cocktail for treating SARS-CoV-2 infections
Zhang, C.; Wang, Y.; Zhu, Y.; Liu, C.; Gu, C.; Xu, S.; Wang, Y.; Zhou, Y.; Wang, Y.; Han, W.; Hong, X.; Yang, Y.; Zhang, X.; Wang, T.; Xu, C.; Hong, Q.; Wang, S.; Zhao, Q.; Qiao, W.; Zang, J.; Kong, L.; Wang, F.; Wang, H.; Qu, D.; Lavillette, D.; Tang, H.; Deng, Q.; Xie, Y.; Cong, Y.; Huang, Z.
Yan, L.; Ge, J.; Zheng, L.; Zhang, Y.; Gao, Y.; Wang, T.; Huang, Y.; Yang, Y.; Gao, S.; Li, M.; Liu, Z.; Wang, H.; Li, Y.; Chen, Y.; Guddat, L. W.; Wang, Q.; Rao, Z.; Lou, Z.
Xu, C.; Wang, Y.; Liu, C.; Zhang, C.; Han, W.; Hong, X.; Wang, Y.; Hong, Q.; Wang, S.; Zhao, Q.; Wang, Y.; Yang, Y.; Chen, K.; Zheng, W.; Kong, L.; Wang, F.; Zuo, Q.; Huang, Z.; Cong, Y.
They spent 12 years solving a puzzle. It yielded the first COVID-19 vaccines.
Kramer, J.
Mihelc. E. M.; Baker, S. C.; Lanman, J. K.
Yuan, S.; Peng, L.; Park, J. J.; Hu, Y.; Devarkar, S. C.; Dong, M. B.; Shen, Q.; Wu, S.; Chen, S.; Lomakin, I. B.; Xiong, Y.
Zhou, T.; Tsybovsky, Y.; Gorman, J.; Rapp, M.; Cerutti, G.; Chuang, G. -Y.; Katsamba, P. S.; Sampson, J. M.; Schön, A.; Bimela, J.; Boyington, J. C.; Nazzari, A.; Olia, A. S.; Shi, W.; Sastry, M.; Stephens, T.; Stuckey, J.; Teng, I. -T.; Kwong, P. D
De novo design of potent and resilient hACE2 decoys to neutralize SARS-CoV-2
Linsky, T. W.; Vergara, R.; Codina, N.; Nelson, J. W.; Walker, M. J.; Su, W.; Barnes, C. O.; Hsiang, T. -Y.; Esser-Nobis, K.; Yu, K.; Reneer, B.; Hou, Y. J.; Priya, T.; Mitsumoto, M.; Pong, A.; Lau, Y.; Mason, M. L.; Chen, J.; Chen, A.; Berrocal, T.; Peng, H.; Clairmont, N. S.; Castellanos, J.; Lin, Y. -R..; Josephson-Day, A.; Baric, R. S.; Fuller, D. H.; Walkey, C. D.; Ross, T. M.; Swanson, R.; Bjorkman, P. J.; Gale Jr., M.; Blancas-Mejia, L. M.; Yen, H. -L.; Silva, D. -A.
Yao, H.; Sun, Y.; Deng, Y. -Q.; Wang, N.; Tan, Y.; Zhang, N. -N.; Li, X. -F.; Kong, C.; Xu, Y. -P.; Chen, Q.; Cao, T. -S.; Zhao, H.; Yan, X.; Cao, L.; Lv, Z.; Zhu, D.; Feng, R.; Wu, N.; Zhang, W.; Hu, Y.; Chen, K.; Zhang, R. -R.; Lv, Q.; Sun, S.; Zhou, Y.; Yan, R.; Yang, G.; Sun, X.; Liu, C.; Lu, X.; Cheng, L.; Qiu, H.; Huang, X. -Y.; Weng, T.; Shi, D.; Jiang, W.; Shao, J.; Wang, L.; Zhang, J.; Jiang, T.; Lang, G.; Qin, C. -F.; Li, L.; Wang, X.
Structure-based development of human antibody cocktails against SARS-CoV-2
Wang, N.; Sun, Y.; Feng, R.; Wang, Y.; Guo, Y.; Zhang, L.; Deng, Y. -Q.; Wang, L.; Cui, Z.; Cao, L.; Zhang, Y. -J.; Li, W.; Zhu, F. -C.; Qin, C. -F.; Wang, X.
Structural analysis of full-length SARS-CoV-2 spike protein from an advanced vaccine candidate
Bangaru, S.; Ozorowski, G.; Turner, H. L.; Antanasijevic, A.; Huang, D.; Wang, X.; Torres, J. L.; Diedrich, J. K.; Tian, J. -H.; Portnoff, A. D.; Patel, N.; Massare, M. J.; Yates III, J. R.; Nemazee, D.; Paulson, J. C.; Glenn, G.; Smith, G.; Ward, A. B.
Ultrapotent human antibodies protect against SARS-CoV-2 challenge via multiple mechanisms
Tortorici, M. A.; Beltramello, M.; Lempp, F. A.; Pinto, D.; Dang, H. V.; Rosen, L. E.; McCallum, M.; Bowen, J.; Minola, A.; Jaconi, S.; Zatta, F.; De Marco, A.; Guarino, B.; Bianchi, S.; Lauron, E. J.; Tucker, H.; Zhou, J.; Peter, A.; Havenar-Daughton, C.; Wojcechowskyj, J. A.; Case, J. B.; Chen, R. E.; Kaiser, H.; Montiel-Ruiz, M.; Meury, M.; Czudnochowski, N.; Spreafico, R.; Dillen, J.; Ng, C.; Sprugasci, N.; Culap, K.; Benigni, F.; Abdelnabi, R.; Foo, S. -Y. C.; Schmid, M. A.; Cameroni, E.; Riva, A.; Gabrieli, A.; Galli, M.; Pizzuto. M. S.; Neyts, J.; Diamond, M. S.; Virgin, H. W.; Snell, G.; Corti, D.; Fink, K.; Veesler, D.;
Architecture of a SARS-CoV-2 mini replication and transcription complex
Yan, L.; Zhang, Y.; Ge, J.; Zheng, L.; Gao, Y.; Wang, T.; Jia, Z.; Wang, H.; Huang, Y.; Li, M.; Wang, Q.; Ra, Z.; Lou, Z.
SARS-CoV-2 structure and replication characterized by in situ cryo-electron tomography
Klein, S.; Cortese, M.; Winter, S. L.; Wachsmuth-Melm, M.; Neufeldt, C. J.; Cerikan, B.; Stanifer, M. L.; Boulant, S.; Bartenschlager, R.; Chlanda , P.
Du, S.; Cao, Y.; Zhu, Q.; Yu, P.; Qi, F.; Wang, G.; Du, X.; Bao, L.; Deng, W.; Zhu, H.; Liu, J.; Nie, J.; Zheng, Y.; Liang, H.; Liu, R.; Gong, S.; Xu, H.; Yisimayi, A.; Qin, C.
Piccoli, L.; Park, Y. -J.; Tortorici, M. A.; Czudnochowski, N.; Walls, A. C.; Beltramello, M.; Silacci-Fregni, C.; Pinto, D.; Rosen, L. E.; Bowen, J. E.; Acton, O. J.; Jaconi, S.; Guarino, B.; Minola, A.; Zatta, F.; Sprugasci, N.; Bassi, J.; Peter, A.; De Marco, A.; Nix, J. C.; Mele, F.; Jovic, S.; Rodriguez, B. F.; Gupta, S. V.; Jin, F.; Piumatti, G.; Presti, G. L.; Pellanda, A. F.; Biggiogero, M.; Tarkowski, M.; Pizzuto, M. S.; Cameroni, E.; Havenar-Daughton, C.; Smithey, M.; Hong, D.; Lepori, V.; Albanese, E.; Ceschi, A.; Bernasconi, E.; Elzi, L.; Ferrari, P.; Garzoni, C.; Riva, A.; Snell, G.; Sallusto, F.; Fink, K.; Virgin, H. W.; Lanzavecchia, A.; Corti, D.; Veesler, D.
Guo, L.; Bi, W.; Wang, X.; Xu, W.; Yan, R.; Zhang, Y.; Zhao, K.; Li, Y.; Zhang, M.; Cai, X.; Jiang, S.; Xie, Y.; Zhou, Q.; Lu, L.; Dang, B.
Free fatty acid binding pocket in the locked structure of SARS-CoV-2 spike protein
Toelzer, C.; Gupta, K.; Yadav, S. K. N.: Borucu, U.; Davidson, A. D.; Williamson, M. K.; Shoemark, D. K.; Garzoni, F.; Staufer, O.; Milligan, R.; Capin, J.; Mulholland, A. J.; Spatz, J.; Fitzgerald, D.; Berger, I.; Schaffitzel, C.
An ultrapotent synthetic nanobody neutralizes SARS-CoV-2 by stabilizing inactive spike
Schoof, M.; Faust, B.; Saunders, R. A.; Sangwan, S.; Rezelj, V.; Hoppe, N.; Boone, M.; Billesbølle, C. B.; Puchades, C.; Azumaya, C. M.; Kratochvil, H. T.; Zimanyi, M.; Deshpande, I.; Liang, J.; Dickinson, S.; Nguyen, H. C.; Chio, C. M.; Merz, G. E.; Thompson, M. C.; Diwanji, D.; Schaefer, K.; Anand, A. A.; Dobzinski, N.; Zha, B. S.; Simoneau, C. R.; Leon, K.; White, K. M.; Chio, U. S.; Gupta, M.; Jin, M.; Li, F.; Liu, Y.; Zhang, K.; Bulkley, D.; Sun, M.; Smith, A. M.; Rizo, A. N.; Moss, F.; Brilot, A. F.; Pourmal, S.; Trenker, R.; Pospiech, T.; Gupta, S.; Barsi-Rhyne, B.; Belyy, V.; Barile-Hill, A. W.; Nock, S.; Liu, Y.; Krogan, N. J.; Ralston, C. Y.; Swaney, D. L.; García-Sastre, A.; Ott, M.; Vignuzzi, M.; QCRG Structural Biology Consortium; Walter, P.; Manglik, A.
Custódio, T. F.; Das, H.; Sheward, D. J.; Hanke, L.; Pazicky, S.; Pieprzyk, J.; Sorgenfrei, M.; Schroer, M. A.; Gruzinov, A. Y.; Jeffries, C. M.; Graewert, M. A.; Svergun, D. I.; Dobrev, N.; Remans, K.; Seeger, M. A.; McInerney, G. M.; Murrell, B.; Hällberg, B. M.; Löw, C.
The architecture of inactivated SARS-CoV-2 with postfusion spikes revealed by cryo-EM and cryo-ET
Liu, C.; Mendonça, L.; Yang, Y.; Gao, Y.; Shen, C.; Liu, J.; Ni, T.; Ju, B.; Liu, C.; Tang, X.; Wei, J.; Ma, X.; Zhu, Y.; Liu, W.; Xu, S.; Liu, Y.; Yuan, J.; Wu, J.; Liu, Z.; Zhang, Z.; Liu, L.; Wang, P.; Zhang, P.
Zhou, T.; Teng, I. -T.; Olia, A. S.; Cerutti, G.; Gorman, J.; Nazzari, A.; Shi, W.; Tsybovsky, Y.; Wang, L.; Wang, S.; Zhang, B.; Zhang, Y.; Katsamba, P. S.; Petrova, Y.; Banach, B. B.; Fahad, A. S.; Liu, L.; Lopez Acevedo, S. N.; Madan, B.; de Souza, M. O.; Pan, X.; Wang, P.; Wolfe, J. R.; Yin, M.; Ho, D. D.; Phung, E.; DiPiazza, A.; Chang, L. A.; Abiona, O. M.; Corbett, K. S.; DeKosky, B. J.; Graham, B. S.; Mascola, J. R.; Misasi, J.; Ruckwardt, T.; Sullivan, N. J.; Shapiro, L.; Kwong, P. D.
De novo design of picomolar SARS-CoV-2 miniprotein inhibitors
Cao, L.; Goreshnik, I.; Coventry, B.; Case, J. B.; Miller, L.; Kozodoy, L.; Chen, R. R.; Carter, L.; Walls, A. C.; Park, Y. -J.; Strauch, E. -M.; Stewart, L.; Diamond, M. S.; Veesler, D.; Baker, D.
SARS-CoV-2 neutralizing antibody structures inform therapeutic strategies
Barnes, C. O.; Jette, C. A.; Abernathy, M. E.; Dam, K. -M., A.; Esswein, S. R.; Gristick, H. B.; Malyutin, A. G.; Sharaf, N. G.; Huey-Tubman, K. E.; Lee, Y. E.; Robbiani, D. F.; Nussenzweig, M. C.; West Jr., A. P.; Bjorkman, P. J.
In situ structural analysis of SARS-CoV-2 spike reveals flexibility mediated by three hinges
Turonova, B.; Sikora, M.; Schurmann, C.; Hagen, W. J. H.; Welsch, S.; Blanc, F. E. C.; von Bulow, S.; Gecht, M.; Bagola, K.; Horner, C.; van Zandbergen, G.; Landry, J.; de Azevedo, N. T. D.; Mosalaganti, S.; Schwarz, A.; Covino, R.; Muhlebach, M. D.; Hummer, G.; Locker, J, K,; Beck, M.
InsteaDMatic: Towards cross-platform automated continuous rotation electron diffraction
Roslova, M.; Smeets, S.; Wang, B.; Thersleff, T.; Xu, H.; Zou, X.
Broad host range of SARS-CoV-2 and the molecular basis for SARS-CoV-2 binding to cat ACE2
Wu, L.; Chen, Q.; Liu, K.; Wang, J.; Han, P.; Zhang, Y.; Hu, Y.; Meng, Y.; Pan, X.; Qiao, C.; Tian, S.; Du, P.; Song, H.; Shi, W.; Qi, J.; Wang, H. -W.; Yan, J.; Gao, G. F.; Wang, Q.
Structure-based design of prefusion-stabilized SARS-CoV-2 spikes
Hseih, C. -L.; Goldsmith, J. A.; Schaub, J. M.; Divenere, A. M.; Kuo, H. -C.; Javanmardi, K.; Le, K. C.; Wrapp, D.; Lee, A. G.; Liu, Y.; Chou, C. -W.; Byrne, P. O.; Hjorth, C. K.; Johnson, N. V.; Ludes-Meyers, J.; Nguyen, A. W.; Park, J.; Wang, N.; Amengor, D.; Lavinder, J. J.; Ippolito, G. C.; Maynard, J. A.; Finkelstein, I. J.; McLellan, J. S.
Structural basis for neutralization of SARS-CoV-2 and SARS-CoV by a potent therapeutic antibody
Lv, Z.; Deng, Y. -Q.; Ye, Q.; Cao, L.; Sun, C. -Y.; Fan, C.; Huang, W.; Sun, S.; Sun, Y.; Zhu, L.; Chen, Q.; Wang, N.; Nie, J.; Cui, Z.; Zhu, D.; Shaw, N.; Li, X. -F.; Li, Q.; Xie, L.; Wang, Y.; Rao, Z.; Qin, C. -F.; Wang, X.
Receptor binding and priming of the spike protein of SARS-CoV-2 for membrane fusion
Benton, D. J.; Wrobel, A. G.; Xu, P.; Roustan, C.; Martin, S. R.; Rosenthal, P. B.; Skehel, J. J.; Gamblin, S. J.
Chen, J.; Malone, B.; Llewellyn, E.; Grasso, M.; Shelton, P. M. M.; Olinares, P. D. B.; Maruthi, K.; Eng, E. T.; Vatandaslar, H.; Chait, B. T.; Kapoor, T. M.; Darst, S. A.; Campbell, E. A.
SARS-CoV-2 Nsp1 binds the ribosomal mRMA channel to inhibit translation
Schubert, K.; Karousis, E. D.; Jomaa, A.; Scaiola, A.; Echeverria, B.; Gurzeler, L. -A.; Leibundgut, M.; Thiel, V.; Mühlemann, O.; Ban, N.
Neutralization of SARS-CoV-2 by destruction of the prefusion spike
Huo, J.; Zhao, Y.; Ren, J.; Zhou, D.; Duyvesteyn, H. M. E.; Ginn, H. M.; Carrique, L.; Malinauskas, T.; Ruza, R. R.; Shah, P. N. M.; Tan, T. K.; Rijal, P.; Coombes, N.; Bewley, K. R.; Tree, J. A.; Radecke, J.; Paterson, N. G.; Supasa, P.; Mongkolsapaya, J.; Screaton, G. R.; Carroll, M.; Townsend, A.; Fry, E. E.; Owens, R. J.; Stuart, D. I.
An alpaca nanobody neutralizes SARS-CoV-2 by blocking receptor interaction
Hanke, L.; Perez, L. V.; Sheward, D. J.; Das, H.; Schulte, T.; Moliner-Morro, A.; Corcoran, M.; Achour, A.; Hedestam, G. B.; Hällberg, B. M.; Murrell, B.; McInerney, G. M.
Structural basis for translational shutdown and immune evasion by the Nsp1 protein of SARS-CoV-2
Thoms, M.; Buschauer, R.; Ameismeier, M.; Loepke, L.; Denk, T.; Hirschenberger, M.; Kratzat, H.; Hayn, M.; Mackens-Kiani, T.; Cheng, J.; Straub, J. H.; Sturzel, C. M.; Frohlich, T.; Berninghausen, O.; Becker, T.; Kirchhoff, F.; Sparrer, K. M.; Beckmann, R.
Molecular architecture of the SARS-CoV-2 virus
Yao, H.; Song, Y.; Chen, Y.; Wu, N.; Xu, J.; Sun, C.; Zhang, J.; Weng, T.; Zhang, Z.; Wu, Z.; Cheng, L.; Shi, D.; Lu, X.; Lei, J.; Crispin, M.; Shi, Y.; Li, L.; Li, S.
Studies in humanized mice and convalescent humans yield a SARS-CoV-2 antibody cocktail
Hansen, J.; Baum, A.; Pascal, K. E.; Russo, V.; Giordano, S.; Wloga, E.; Fulton, B. O.; Yan, Y.; Koon, K.; Patel, K.; Chung, K. M.; ermann, A.; Ullman, E.; Cruz, J.; Rafique, A.; Huang, T.; Fairhurst, J.; Libertiny, C.; Malbec, M.; Lee, W. -Y.; Welsh, R.; Farr, G.; Pennington, S.; Deshpande, D.; Cheng, J.; Watty, A.; Bouffard, P.; Babb, R.; Levenkova, N.; Chen, C.; Zhang, B.; Hernandez, A. R.; Saotome, K.; Zhou, Y.; Franklin, M.; Sivapalasingam, S.; Lye, D. C.; Weston, S.; Lopgue, J.; Haupt, R.; Frieman, M.; Chen., G.; Olson, W.; Murphy, A. J.; Stahl, N.; Yancopoulos, G. D.; Kyratsous, C. A.
Barnes, C. O.; West Jr., A. P.; Huey-Tubman, K. E.; Hoffmann, M. A. G.; Sharaf, N. G.; Hoffman, P. R.; Koranda, N.; Gristick, H. B.; Gaebler, C.; Muecksch, F.; Lorenzi, J. C. C.; Finkin, S.; Hägglöf, T.; Hurley, A.; Millard, K. G.; Weisblum, Y.; Schmidt, F.; Hatziioannou, T.; Bieniasz, P. D.; Caskey, M.; Robbiani, D. F.; Nussenzweig, M. C.; Bjorkman. P. J.
A neutralizing human antibody binds to the N-terminal domain of the spike protein of SARS-CoV-2
Chi, X.; Yan, R.; Zhang, G.; Zhang, Z.; Hao, M.; Zhang, Z.; Fan, P.; Dong, X.; Yang, Y.; Chen, Z.; Guo, Y.; Zhang, J.; Li, Y.; Song, X.; Cheng, Y.; Xia, L.; Fu, L.; Hou, L.; Xu, J.; Yu, X.; Li, J.; Zhou, Q.; Chen, W.
Structural basis for the neutralization of SARS-CoV-2 by an antibody from a convalescent patient
Zhou, D.; Duyvesteyn, H. M. E.; Chen, C. -P.; Huang, C. -G.; Chen, T. -H.; Shih, S. -R.; Lin, Y. -C.; Cheng, C. -Y.; Cheng, S. -H.; Huang, Y. -C.; Lin, T. -Y.; Ma, C.; Huo, J.; Carrique, L.; Malinauskas, T.; Ruza, R. R.; Shah, P. N. M.; Tan, T. K.; Rijal, P.; Donat, R. F.; Godwin, K.; Buttigieg, K. R.; Tree, J. A.; Radecke, J.; Paterson, N. G.; Supasa, P.; Mongkolsapaya, J.; Screaton, G. R.; Carroll, M. W.; Gilbert-Jaramillo, J.; Knight, M. L.; James, W.; Owens, R. J.; Naismith, J. H.; Townsend, A. R.; Fry, E. E.; Zhao, Y.; Ren, J.; Stuart, D. I.; Huang, K. -Y. A.
A thermostable, closed SARS-CoV-2 spike protein trimer
Xiong, X.; Qu, K.; Ciazynska, K. A.; Hosmillo, M.; Carter, A. P.; Ebrahimi, S.; Ke, Z.; Scheres, S. H. W.; Bergamaschi, L.; Grice, G. L.; Zhang, Y.; The CITIID-NIHR COVID-19 BioResource Collaboration; Nathan, J. A.; Baker, S.; James, L. C.; Baxendale, H. E.; Goodfellow, I.; Doffinger, R.; Briggs, J. A. G.
Structural basis for RNA replication by the SARS-CoV-2 polymerase
Wang, Q.; Wu2, J.; Wang, H.; Gao, Y.; Liu, Q.; Mu, A.; Ji, W.; Yan, L.; Zhu, Y.; Zhu, C.; Fang, X.; Yang, X.; Huang, Y.; Gao, H.; Liu, F.; Ge, J.; Sun, Q.; Yang, X.; Xu, W.; Liu, Z.; Yang, H.; Lou, Z.; Jiang, B.; Guddat, L. W.; Gong, P.; Rao, Z.
Controlling the SARS-CoV-2 spike glycoprotein conformation
Henderson, R.; Edwards, R. J.; Mansouri, K.; Janowska, K.; Stalls, V.; Gobeil, S. M. C.; Kopp, M.; Li, D.; Parks, R.; Hsu, A. L.; Borgnia, M. J.; Haynes, B. F.; Acharya, P.
Cryo-EM analysis of the post-fusion structure of the SARS-CoV spike glycoprotein
Fan, X.; Cao, D.; Kong, L.; Zhang, X.
Potently neutralizing and protective human antibodies against SARS-CoV-2
Zost, S. J.; Gilchuk, P.; Case, J. B.; Binshtein, E.; Chen, R. E.; Nkolola, J. P.; Schäfer, A.; Reidy, J. X.; Trivette, A.; Nargi, R. S.; Sutton, R. E.; Suryadevara, N.; Martinez, D. R.; Williamson, L. E.; Chen, E. C.; Jones, T.; Day, S.; Myers, L.; Hassan, A. O.; Kafai, N. M.; Winkler, E. S.; Fox, J. M.; Shrihari, S.; Mueller, B. K.; Meiler, J.; Chandrashekar, A.; Mercado, N. B.; Steinhardt, J. J.; Ren, K.; Loo, Y. -M.; Kallewaard, N. L.; McCune, B. T.; Keeler, S. P.; Holtzman, M. J.; Barouch, D. H.; Gralinski, L. E.; Baric, R. S.; Thackray, L. B.; Diamond, M. S.; Carnahan, R. H.; Crowe Jr, J. E.
Cao, Y.; Su, B.; Guo, X.; Sun, W.; Deng, Y.; Bao, L.; Zhu, Q.; Zhang, X.; Zheng, Y.; Geng, C.; Chai, X.; He, R.; Li, X.; Lv, Q.; Zhu, H.; Deng, W.; Xu, Y.; Wang, Y.; Qiao, L.; Tan, Y.; Song, L.; Wang, G.; Du, X.; Gao, N.; Liu, J.; Xiao, J.; Su, X. -D.; Du, Z.; Feng, Y.; Qin, C.; Qin, C.; Jin, R.; Xie, X. S.
Structural basis for inhibition of the RNA-dependent RNA polymerase from SARS-CoV-2 by remdesivir
Yin, W.; Mao, C.; Luan, X.; Shen, D. -D.; Shen, Q.; Su, H.; Wang, X.; Zhou, F.; Zhao, W.; Gao, M.; Chang, S.; Xie, Y. -C.; Tian, G.; Jiang, H. -W.; Tao, S. -C.; Shen, J.; Jiang, Y.; Jiang, H.; X.u, Y.; Zhang, S.; Xu, H. E.
Langer, L. M.; Gat, Y.; Bonneau, F.; Conti, E.
Structure of replicating SARS-CoV-2 polymerase
Hillen, H. S.; Kokic, G.; Farnung, L.; Dienemann, C.; Tegunov, D.; Cramer, P.
Structure, function, and antigenicity of the SARS-CoV-2 spike glycoprotein
Walls, A. C.; Park, Y. -J.; Tortorici, M. A.; Wall, A.; McGuire, A. T.; Veesler, D.
Structural basis for the recognition of the SARS-CoV-2 by full-length human ACE2
Yan, R.; Zhang, Y.; Li, Y.; Xia, L.; Guo, Y.; Zhou, Q.
Structure, function and antigenicity of the SARS-CoV-2 spike glycoprotein
Walls, A. C.; Park, Y. -J.; Tortorici, M. A.; Wall, A.; McGuire, A. T.; Veesler, D
Celia, H.; Botos, I.; Ni, X.; Fox, T.; De Val, N.; Lloubes, R.; Jiang, J.; Buchanan, S. K.
Improved applicability and robustness of fast cryo-electron tomography data acquisition
Eisenstein, F.; Danev, R.; Pilhofer, M.
Solving a new R2lox protein structure by microcrystal electron diffraction
Xu, H.; Lebrette, H.; Clabbers, M. T. B.; Zhao, J.; Griese, J. J.; Zou, X.; Hogbom, M.
Surpassing the physical Nyquist limit to produce super-resolution cryo-EM reconstructions
Feathers, J. R.; Spoth, K. A.; Fromme, J. C.
Single particle cryo-EM reconstruction of 52 kDa streptavidin at 3.2 Angstrom resolution
Fan, X.; Wang, J.; Zhang, X.; Yang, Z.; Zhang, J. -C.; Zhao, L.; Peng, H. -L.; Lei, J.; Wang, H. -W.
Fusion-dependent formation of lipid nanoparticles containing macromolecular payloads
Kulkarni, J. A.; Witzigmann, D.; Leung, J.; van der Meel, R.; Zaifman, J.; Darjuan, M. M.; Grisch-Chan, H. M.; Thöny, B.; Tam, Y. Y. C.; Cullis, P. R.
Putti, M.; Stassen, O. M. J. A.; Schotman, M. J. G.; Sahlgren, C. M.; Dankers, P. Y. W.
A 3.8 Å resolution cryo-EM structure of a small protein bound to an imaging scaffold
Liu, Y.; Huynh, D. T.; Yeates, T. O.
In situ structures of polar and lateral flagella revealed by cryo-electron tomography
Zhu, S.; Schniederberend, M.; Zhitnitsky, D.; Jain, R.; Galán, J. E.; Kazmierczak, B. I.; Liu, J.
Refinement and analysis of the mature Zika virus cryo-EM structure at 3.1 Å resolution
Sevvana, M.; Long, F.; Miller, A. S.; Klose, T.; Buda, G.; Sun, L.; Kuhn, R. J.; Rossmann, M. G.
Xu, H. Lebrette, H.; Yang, T.; Hovmoller, S.; Hogbom, M.; Zou, X.
Cryo-EM structure of 5-HT3A receptor in its resting conformation
Basak, S.; Gicheru, Y.; Samanta, A.; Molugu, S. K.; Huang, W.; la de Fuente, M.; Hughes, T.; Taylor, D. J.; Nieman, M. T.; Moiseenkova-Bell, V.; Chakrapani, S.
Related materials
Nyquist frequency
Dose fractionation and motion correction
Improving DQE with counting and super-resolution
| Modulation transfer function (MTF) curves | ||
|---|---|---|
| 200 kV | 300 kV | |
|
K3 |
CDS | CDS |
| Standard | Standard | |
| K2 | Standard | Standard |